14. beta_profile_gene_centered.py¶
14.1. Description¶
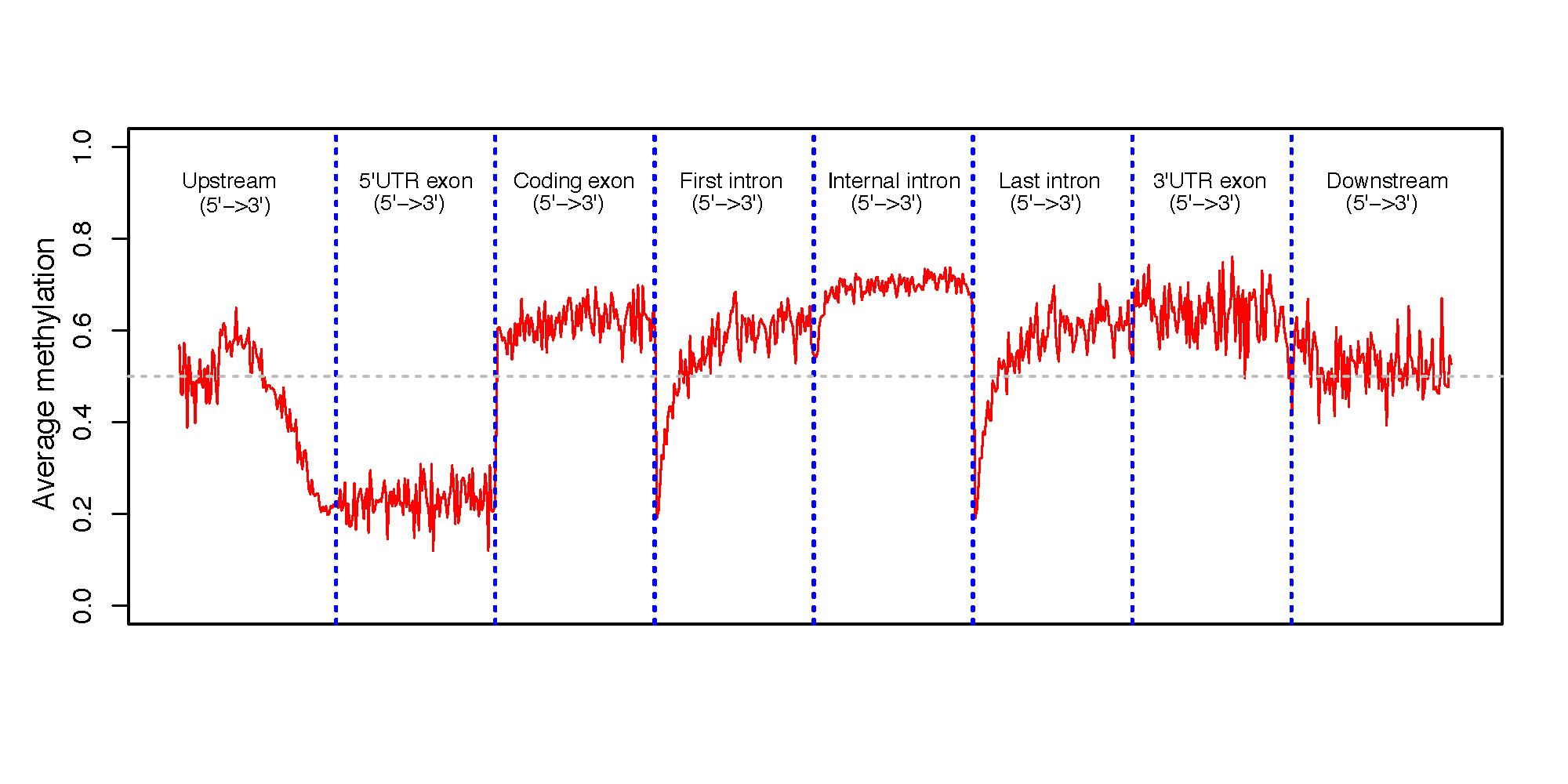
This program calculates the methylation profile (i.e., average beta value) for genomic regions around genes. These genomic regions include:
- 5’UTR exon
- CDS exon
- 3’UTR exon,
- first intron
- internal intron
- last intron
- up-stream intergenic
- down-stream intergenic
Example of input (BED6+)
chr22 44021512 44021513 cg24055475 0.9231 -
chr13 111568382 111568383 cg06540715 0.1071 +
chr20 44033594 44033595 cg21482942 0.6122 -
14.2. Options¶
--version show program’s version number and exit -h, --help show this help message and exit -i INPUT_FILE, --input_file=INPUT_FILE BED6+ file specifying the C position. This BED file should have at least 6 columns (Chrom, ChromStart, ChromeEnd, Name, Beta_value, Strand). BED6+ file can be a regular text file or compressed file (.gz, .bz2). -r GENE_FILE, --refgene=GENE_FILE Reference gene model in standard BED12 format (https://genome.ucsc.edu/FAQ/FAQformat.html#format1). “Strand” column must exist in order to decide 5’ and 3’ UTRs, up- and down-stream intergenic regions. -d DOWNSTREAM_SIZE, --downstream=DOWNSTREAM_SIZE Size of down-stream genomic region added to gene. default=2000 (bp) -u UPSTREAM_SIZE, --upstream=UPSTREAM_SIZE Size of up-stream genomic region added to gene. default=2000 (bp) -o OUT_FILE, --output=OUT_FILE The prefix of the output file.
14.3. Command¶
$beta_profile_gene_centered.py -i test_02.bed6.gz -r hg19.RefSeq.union.bed.gz -o gene_profile