11. beta_UMAP.py¶
11.1. Description¶
This program performs UMAP (Uniform Manifold Approximation and Projection) non-linear dimension reduction.
Example of input data file
ID Sample_01 Sample_02 Sample_03 Sample_04
cg_001 0.831035 0.878022 0.794427 0.880911
cg_002 0.249544 0.209949 0.234294 0.236680
cg_003 0.845065 0.843957 0.840184 0.824286
...
Example of input group file
Sample,Group
Sample_01,normal
Sample_02,normal
Sample_03,tumor
Sample_04,tumo
...
Notes
- Rows with missing values will be removed
- Beta values will be standardized into z scores
- Only the first two components will be visualized
- Options:
--version show program’s version number and exit -h, --help show this help message and exit -i INPUT_FILE, --input_file=INPUT_FILE Tab-separated data frame file containing beta values with the 1st row containing sample IDs and the 1st column containing CpG IDs. -g GROUP_FILE, --group=GROUP_FILE Comma-separated group file defining the biological groups of each sample. Different groups will be colored differently in the 2-dimensional plot. Supports a maximum of 20 groups. -n N_COMPONENTS, --ncomponent=N_COMPONENTS Number of components. default=2 --nneighbors=N_NEIGHBORS This parameter controls the size of the local neighborhood UMAP will look at when attempting to learn the manifold structure of the data. Low values of ‘–nneighbors’ will force UMAP to concentrate on local structure, while large values will push UMAP to look at larger neighborhoods of each point when estimating the manifold structure of the data. Choose a value from [2, 200]. default=15 --min-dist=MIN_DISTANCE This parameter controls how tightly UMAP is allowed to pack points together. Choose a value from [0, 1). default=0.2 -l, --label If True, sample ids will be added underneath the data point. default=False -c PLOT_CHAR, --char=PLOT_CHAR Ploting character: 1 = ‘dot’, 2 = ‘circle’. default=1 -a PLOT_ALPHA, --alpha=PLOT_ALPHA Opacity of dots. default=0.5 -x LEGEND_LOCATION, --loc=LEGEND_LOCATION Location of legend panel: 1 = ‘topright’, 2 = ‘bottomright’, 3 = ‘bottomleft’, 4 = ‘topleft’. default=1 -o OUT_FILE, --output=OUT_FILE The prefix of the output file.
11.2. Input files (examples)¶
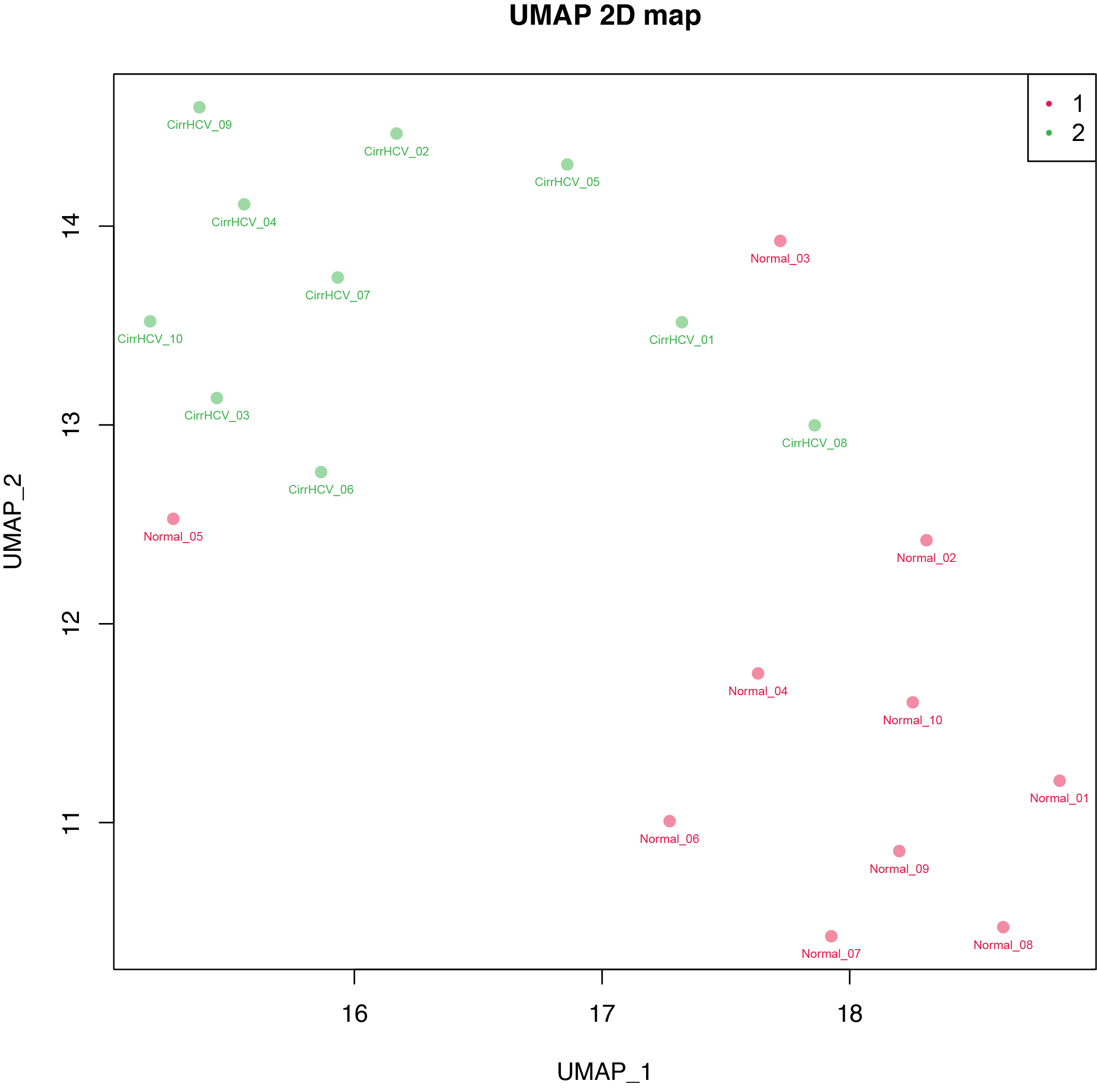
11.3. Command¶
$beta_UMAP.py -i cirrHCV_vs_normal.data.tsv -g cirrHCV_vs_normal.grp.csv -o cirrHCV_vs_normal -l